Editor's Choice: Defining principles for mobile apps and platforms development in citizen science
By: Ulrike Sturm, Sven Schade, Luigi Ceccaroni, Margaret Gold, Christopher Kyba, Bernat Claramunt, Muki Haklay, Dick Kasperowski, Alexandra Albert, Jaume Piera, Jonathan Brier, Christopher Kullenberg, Soledad Luna
Editor's Choice: The focus of this paper is a summary of two workshops held to better understand the design and development principles for mobile or web-based citizen science platforms. The recommendations from the working groups are well worth reading - even if you think you know everything about developing citizen science projects! - LFF Abstract: Apps for mobile devices and web-based platforms are increasingly used in citizen science projects. While…
Read More
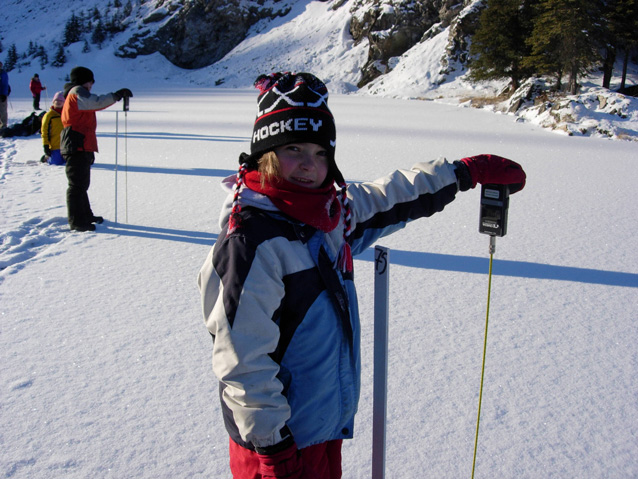
Editor's Choice: Contrasting the Views and Actions of Data Collectors and Data Consumers in…
By: Caren B. Cooper, Lincoln R. Larson, Kathleen Krafte Holland, Rebecca A. Gibson, David J. Farnham, Diana Y. Hsueh, Patricia J. Culligan, Wade R. McGillis
Editor's Choice: This paper points out that there is an oft-overlooked category of people interacting with citizen science projects - people who do not contribute to the data collection, but who nonetheless consume data and information related to the project. The questions are then posed - do these data consumers share the same characteristics as the data producers; and do they actually impact project outcomes? If you have anything to…
Read More
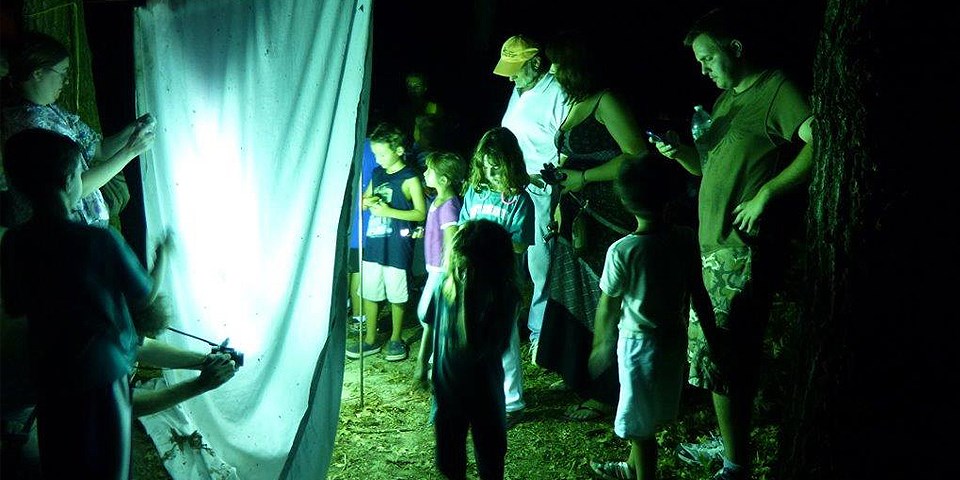
Happy New Year from Citizen Science Today
By: CST Editors, LFF & AWA
What a year we saw with 2017! Staying focused on the positive, it was a great year for citizen science. Here are just a few call-outs but all citizen scientists should immediately pat themselves on the back! We CAN do it and have fun at the same time. This year, the Citizen Science Association hosted its annual meeting right here in the Twin Cities (from where we are writing) with…
Read More
Editor's Choice: Is ‘grassroots’ citizen science a front for big business?
By: Philip Mirowski
Editor's Choice: This is a rather provocative essay whose arguments are completely specious in many instances, and completely wrong in others, but has enough seeds of truth that it is worth pondering and acknowledging, and then considering much-needed rebuttals. --LFF-- Excerpt: The very label ‘citizen science’ (as opposed to, say, ‘amateur’ or ‘extramural’) carries the unsubtle suggestion that science should be a participatory democracy, not an unpalatable, autocratic regime. Proponents claim…
Read More